Researchers from Rockefeller University used a C-Trap® optical tweezers – fluorescence microscopy system to observe the behavior of the main component of the replisome – the eukaryotic CMG helicase – during DNA replication. The study offers exciting new insights into the interactions, processes, and structural plasticity associated with CMG during DNA binding and replication and explains how replication forks can continue past DNA lesions and start again after stalling.
These findings were published in the journal Cell in the article “Replication Fork Activation Is Enabled by a Single-Stranded DNA Gate in CMG Helicase”. Congratulations to all the authors!
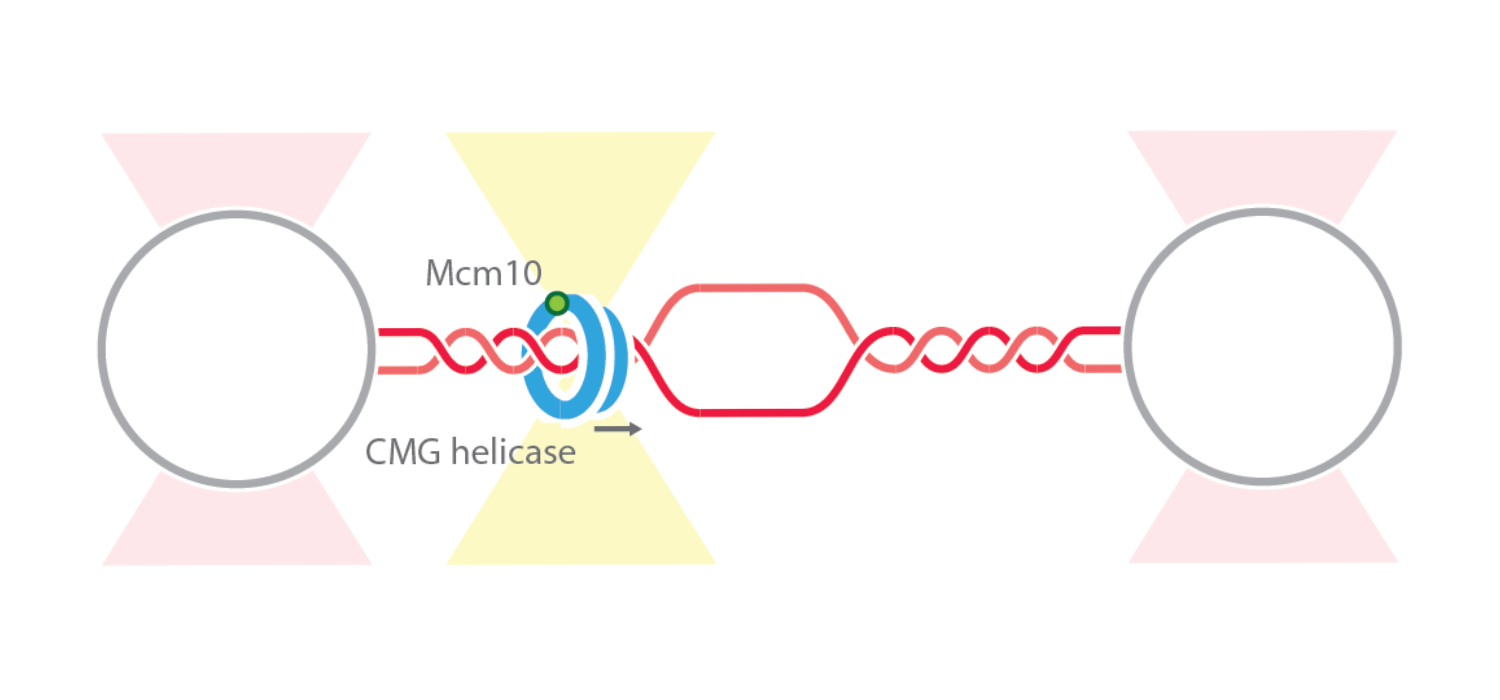
Wasserman et al. analyzed the properties of fluorescently-labeled CMG during DNA replication. For this, they used optical tweezers to tether a double-stranded molecule between two beads and create DNA forks in situ by force manipulation. They then visualized the behavior of single CMG complexes using fluorescence imaging while subjecting the replisome assembly to different reaction conditions by utilizing the laminar flow microfluidics.
By applying high tension to the double-stranded DNA molecule, the investigators could establish regions of single-stranded DNA (so-called DNA bubbles) and study the affinity of CMG to either of the two DNA conformations. Also, by fluorescently labeling additional components, they could further analyze CMG dynamics upon DNA association. For example, they found that CMG forms a complex with replisome protein Mcm10 and binds DNA at fork junctions (between double-stranded and single-stranded DNA). On top of that, they showed that the complex induced active replication at the fork junction.
Importantly, through DNA stretching, the team found that the CMG/Mcm10 complex uncoupled from the polymerase and could transition from single-stranded to double-stranded DNA upon replication stress by employing a gate in its ring. In other words, the CMG helicase is maintained on the DNA – albeit on a double-stranded region – while vacating a stalled fork, and re-enters the fork once the repair is completed.
The C-Trap® instrument used in this study is commercially available by LUMICKS. Are you interested in using them in your research? Please feel free to contact us for a demo or quote.