A recent publication by Dai et al. in Nature Communications presents an optical tweezers-coupled Raman spectroscopy platform providing tuneable and reproducible surface-enhanced Raman spectroscopy (SERS) measurements. By combining a Raman microscope with the dual-trap optical tweezers and microfluidic flow chamber of the m-Trap®, the scientists were able to characterize three different protein structures, including transient species of α-synuclein.
The combined platform enables the measurement of analytes at physiological concentrations of around several micro-molars and can be particularly useful for the analysis of intrinsically disordered proteins (IDPs). IDPs lack stable secondary and tertiary structures and can stabilize in aggregates, which are known to cause neurodegenerative diseases. Due to the transient nature of IDP conformations and low population of each conformational state, the structural characterization of IDPs is challenging and requires new methodological approaches.
By facilitating easy detection of proteins at low concentrations and enabling highly controllable, sensitive measurements, the combination of the m-Trap® and Raman spectrometer can contribute to resolving the structures and dynamics of heterogeneous biological systems like IDPs without perturbating their native states.
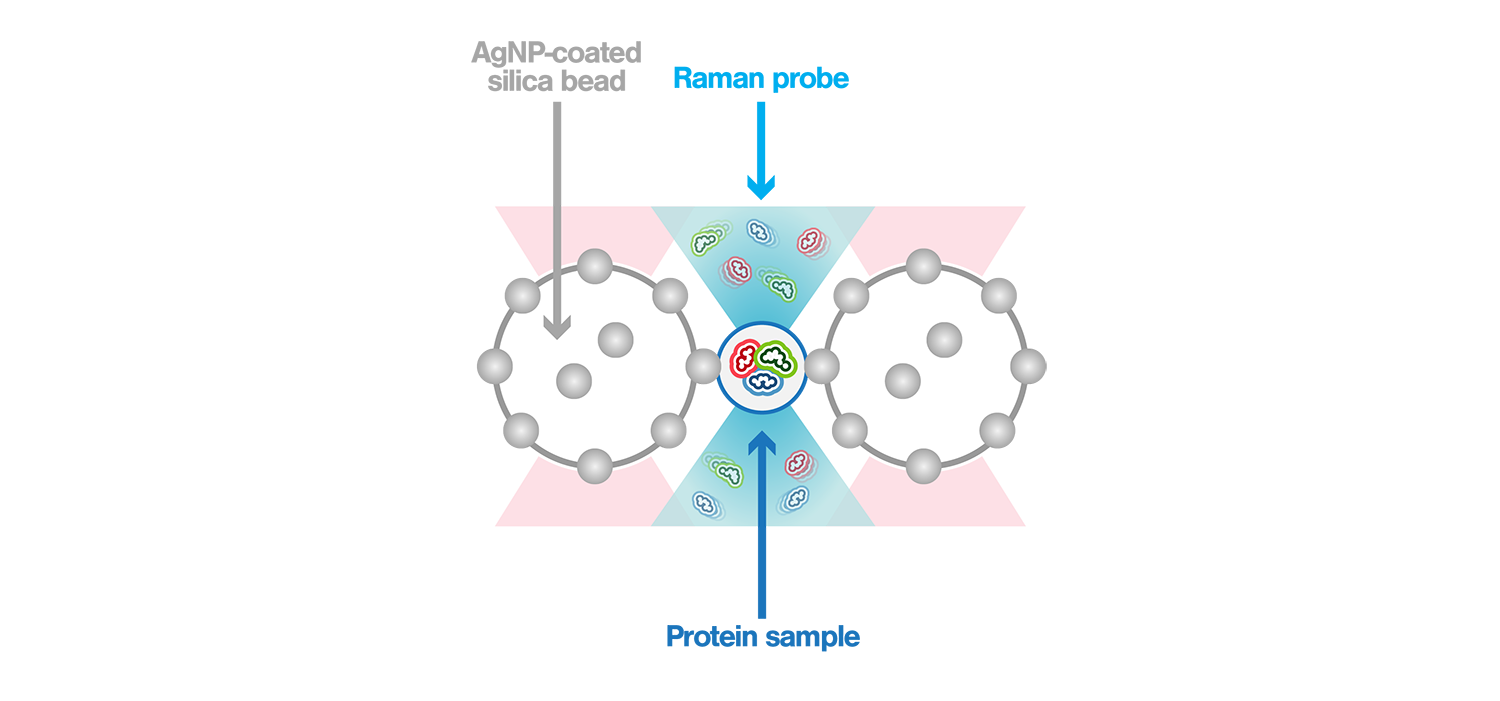
Surface-enhanced Raman spectroscopy (SERS) is a powerful method used to characterize biomolecules in aqueous solutions. Its power comes from the high enhancement of the Raman signal that is generated when the sample is measured within the “hotspot”, a strong electromagnetic field confined between plasmonic nanostructures. Using the m-Trap®–coupled Raman spectrometer, the scientists generated the hotspot between two optically trapped silver nanoparticle (AgNP) – coated silica beads. This allowed to precisely control the hotspot in 10 nm steps and directly observe its location through brightfield imaging, thus generating a dynamic SERS window with real-time controllable hotspots. This dynamic SERS window is placed in the m-Trap® microfluidic flow chamber to detect passing-by proteins, enabling characterization of their structure without perturbation of their native states.
Employing this method, the scientists performed up to 200 reproducible measurements, analyzing the chemical structures of 100 nM hemoglobin, 1 µM lysozyme, and the transient conformations of 1 µM α-synuclein.
This novel approach offers high tunability and reproducibility of SERS enhancement, both highly relevant for gathering reliable data and so far not achieved by other methods. This was facilitated by the possibility of the m-Trap® allowing an easy integration of additional imaging modules, like Raman spectroscopy, into its provided optical flexport. This combined system allows the structural analysis of proteins at physiological concentrations without perturbating their native states. Notably, it enables the characterization of transient protein conformations, and can contribute to the understanding of intrinsically disordered proteins (IDPs) and their transition into pathology-inducing aggregates.
To find out more, read the full article titled “Optical tweezers-controlled hotspot for sensitive and reproducible surface-enhanced Raman spectroscopy characterization of native protein structures” published in Nature Communications.
The versatility of the m-Trap® platform allows to add a wide variety of method modules. Click here to learn more about the product.