In a recent Science Advances publication, the group of Sander J. Tans from the AMOLF Institute, The Netherlands, demonstrate for the first time, at the single molecule level, how the molecular chaperone GroEL-ES modulates protein chain collapse and stimulates protein folding. Using LUMICKS’ C-Trap®, they were able to follow and visualize single protein chains as they entered the cavity of a GroEL chaperone, discovering how protein folding can be accelerated.
Protein compaction plays a crucial role in molecular processes such as protein folding, aggregation and phase separation. Unfolded proteins are known to collapse autonomously into compact structure, but whether protein collapse can be controlled by the cellular machinery remains to be elucidated. GroEL-GroES is a well-studied chaperone, however, the mechanism by which it stimulates protein folding remains unclear.
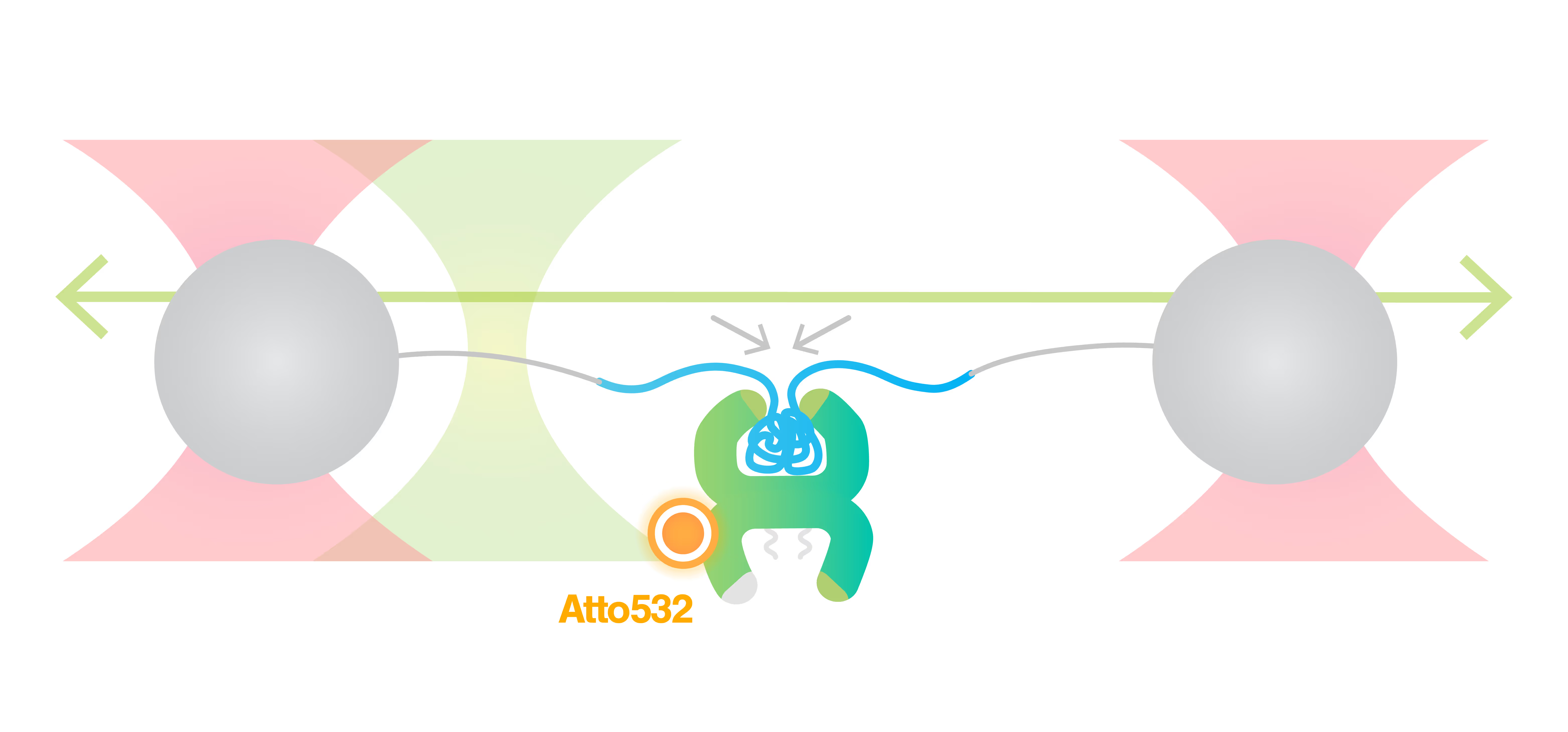
The combination of optical tweezers, multichannel microfluidics and fluorescent microscopy offered by the C-Trap allowed Naqvi and colleagues to monitor real-time interactions between the maltose-binding protein (MBP) and the chaperone GroEL-ES. By attaching MBP to polystyrene beads and manipulating the distance between the protein ends by moving the laser traps to displace the beads and stretch the protein, they were able to monitor substrate protein folding behavior. Surprisingly, they observed that GroEL-ES employs a folding acceleration mechanism to accelerate MBP folding. When adding GroEL to the solution alone, the two ends of MBP started moving closer together and the protein started collapsing into the GroEL cavity. Then the protein started folding rapidly. Adding GroES, the “lid” on the cavity of GroEL, demonstrated that GroES is not required to accelerate protein folding, but is critical in the generation of fully folded proteins and their ejection from the GroEL cavity. Naqvi and coworkers went on to monitor the movements of fluorescently labelled GroEL while manipulating MBP using optical traps. Fluorescent GroEL binds to the substrate protein right before the folding event, confirming that GroEL is, indeed, responsible for it. This integrated technique also allowed the scientists to verify the formation of a ternary complex GroEL-ES-MBP.
“The optical trap assay detects distinct conformational transitions that occur on the time scale of a few seconds.”
Based on these experiments the authors suggest a model in which GroEL pulls MPB inside its cavity by triggering contractile forces in the substrate, thereby inducing collapse and folding transitions. GroES helps this collapse by binding to the cavity rim.
“Folding acceleration mechanisms have never been shown before for any protein system. This is arguably the most beautiful protein research we have ever done in our laboratory. A highly surprising mechanism, and all observations and control measurements match perfectly,” Tans said.
These first-of-their-kind exciting findings have implications for our understanding of cellular protein control and folding diseases. The notion that protein collapse and folding speeds can be regulated may shine a new light on diseases where protein folding, and aggregation plays an important role. For instance, it offers new avenues to study the regulation of disordered oncogenic c-Myc/Max complex, the phase separated FUS proteins, which play a role in ALS and the aggregating tau protein associated with degenerative brain diseases.
For further information about this study, check out the paper “Protein chain collapse modulation and folding stimulation by GroEL-ES”, published in Science Advances.
Are you interested in using dynamic single-molecule tools like the C-Trap for your research? Do not hesitate to contact us for more information, a demo, or a quote.
Top image adapted from Naqvi et al., Science Advances, 2022.
This blog post includes information and quotes taken from a press release published at Phys.org.