A recent paper published in Nature by the group of Dr. Simon Boulton from the Francis Crick Institute enlightens the role played by the helicase HELQ in the context of DNA repair. The researchers took advantage of the combination of microfluidics, optical tweezers, and fluorescence microscopy of the C-Trap® to visualize the interaction between HELQ and DNA in real time at single-molecule level. Moreover, they could clarify the interplay between HELQ and two other key proteins in the process of DNA repair, i.e. replication protein A (RPA) and RAD51.
DNA double-strand breaks (DSBs) are hazardous lesions, whose erroneous repair can lead to cancer development. HELQ is required for DNA repair, and its deletion in mice has been shown to increase the risk of ovarian and pituitary tumors. Furthermore, HELQ deficiency causes hypersensitivity to anti-tumor agents at the cell level. Although this helicase is known to bind to RPA and the RAD51 paralog BCDX2 complex, the role of this interaction and the exact function played by HELQ in the mechanism of DSB repair remains unknown.
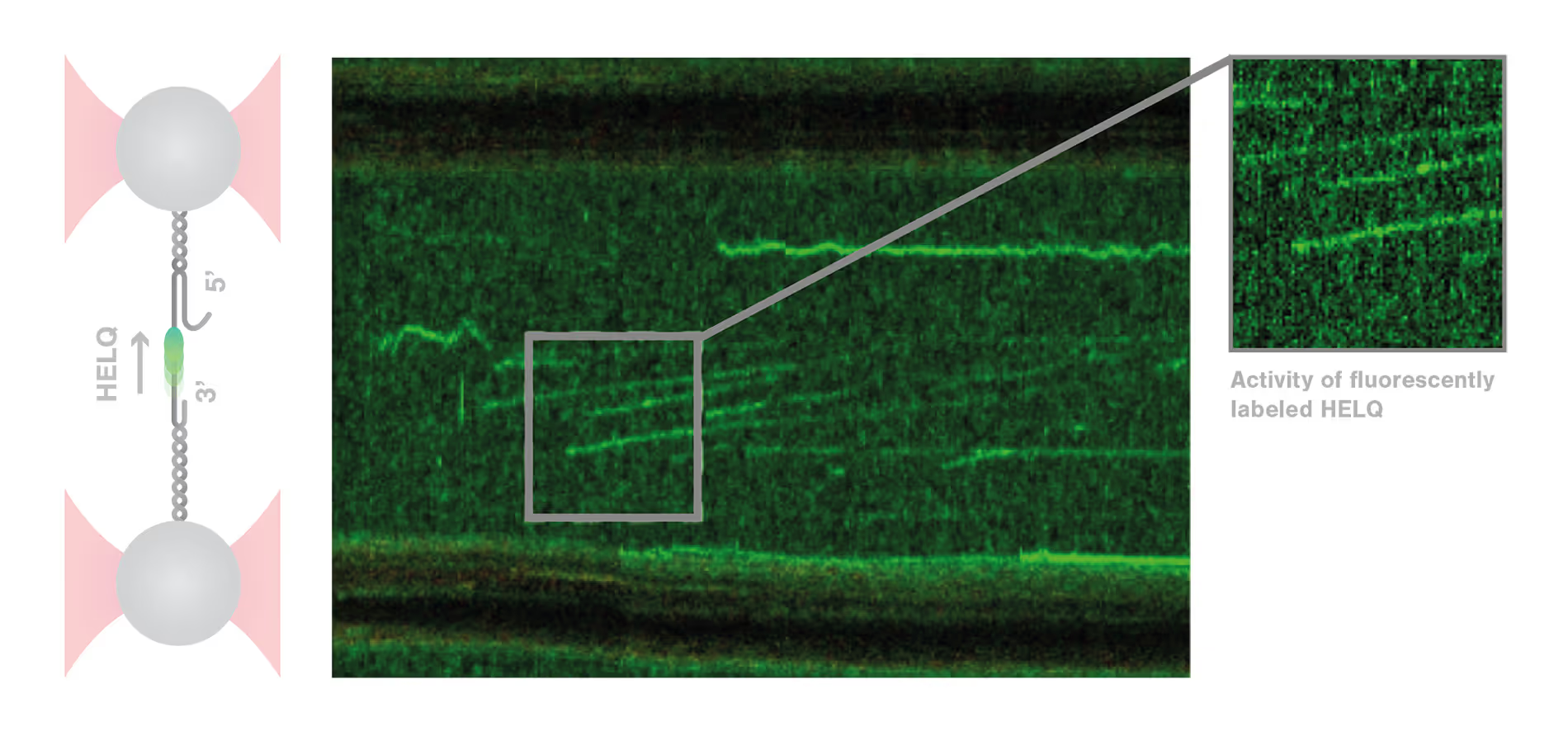
To answer these questions, Roopesh Anand and colleagues used the C-Trap to mimic some crucial steps of DSB repair in vitro and test the influence of different conditions. The scientists tethered and stretched an individual double-stranded (ds) DNA molecule containing a single-stranded (ss) DNA gap between two optically trapped beads. When they added HELQ, the distance between the beads increased, revealing the binding and unwinding of dsDNA by HELQ. Strikingly, when the DNA-HELQ construct was moved into a microfluidics channel containing also RAD51, the unwinding speed of the helicase markedly increased. Moreover, using the confocal microscopy module of the C-Trap, the researchers could directly visualize RAD51 translocating with the helicase on the DNA filaments and follow the unwinding of the DNA in real-time. These results revealed that HELQ forms a complex with RAD51 which unwinds the dsDNA three times faster than HELQ alone.
Besides its well-known unwinding activity, the researchers found that HELQ can also (re)anneal ssDNA strands and that, at low helicase concentrations, the addition of RPA accelerates the annealing reaction. Based on this result, the authors hypothesized that RPA helps to load HELQ onto ssDNA when HELQ is available at a low concentration. HELQ was also shown to efficiently displace the RPA bound to ssDNA thus facilitating the annealing. Finally, the authors demonstrated that, with the help of RPA, ssDNA-binding HELQ can capture another ssDNA in a sequence-independent manner. Such a process was visualized in real-time with the C-Trap using a construct of ssDNA binding both RPA and HELQ and additional short fluorescently-labeled ssDNA sequences. Finally, the authors performed in vivo experiments to prove the physiological role of HELQ in a number of DNA repair processes, such as single-strand annealing and homologous recombination.
This work is highly relevant as it unravels the mechanisms of function of HELQ in DNA DSB repair and its interplay with other crucial proteins in this process. Ultimately, these insights will be relevant to learning more about genetic instability and tumor development. In this study, the scientists used a unique combination of single-molecule technology, biochemistry methods, and in vitro and in vivo experiments to make novel observations and obtain mechanistic insights. To read more about their findings, check their article with the title: “HELQ is a dual-function DSB repair enzyme modulated by RPA and RAD51”, published in Nature.